Many people who decide to commit suicide do not behave too adequately. Someone is constantly in a sullen mood, someone else begins to behave aggressively, but it also happens that a potential suicide does not give himself away. Sometimes it happens that today a person is still having fun with friends, playing guitar and singing, and tomorrow he decides to commit suicide.
According to statistics, about
800,000 people commit suicide every year. Sometimes it happens that the situation, because of which a person decided to die, is not at all hopeless, and if someone spoke to a potential suicide before his death, then the person would simply abandon his idea. Now scientists at Carnegie Mellon University are
developing a neural network that can predict when a person decides to kill himself.
According to scientists, when a person makes such a decision, certain areas of the brain are involved. “Our last job is to identify brain iterations that are associated with suicidal tendencies,” said Marcel Just, one of the representatives of the project. “This gives us the opportunity to open the window to the brain and see how the brain of a potential suicide works.”
In the course of previous research papers, Just and his colleagues created computer models of the brain for mapping individual sites responsible for certain thoughts and moods. In particular, they were able to identify the signature of sadness, shame, anger and other negative emotions.
The group of volunteers who took part in the project was small - only 34 people. Of these, 17 patients had suicidal tendencies (about half of the participants in this group attempted to commit suicide earlier) and 17 people were completely normal, without suicidal tendencies.
Next, participants underwent an operation such as functional magnetic resonance imaging. During the scan, the volunteer showed 10 cards with words related to suicide (despair, hopelessness, lifelessness), 10 cards with positive words (for example, “carefree”) and 10 cards with just words with negative emotional overtones (“problem”).
After all the volunteers went through the scanning procedure, the scientists chose six terms with the maximum emotional response of the volunteers - “death”, “cruelty”, “problem”, “carelessness”, “good”, “praise”. There were also five areas of the brain, which work for the "suicides" during the demonstration of cards was different from the work of "normal" people.
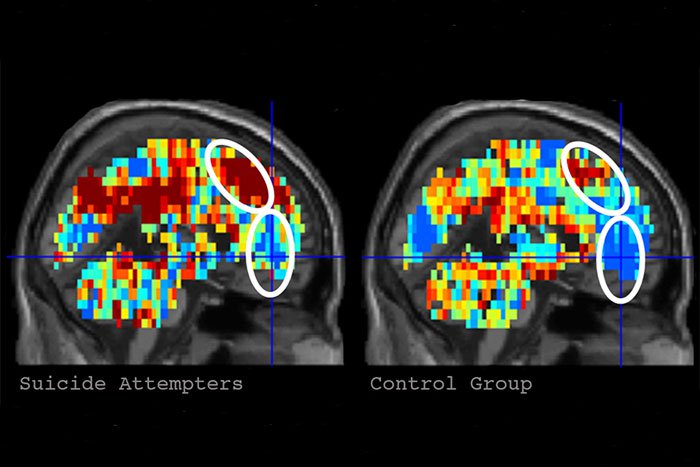
Using this very small set of data, experts began to train the neural network to respond to a similar picture of the brain. At the end of the learning process, the system determined the suicidal tendency in 9 cases out of 10. In addition, the system learned to distinguish those potential suicides who had already tried to commit suicide from those who had never tried to do so. The accuracy of identification in this case was 94%. As it turned out, potential suicides give the maximum emotional response to the words “death”, “lifeless” and “carefree”.
“In the future, we will need a larger sample in order to teach the system to accurately determine suicidal behavior and to give physicians the ability to identify people with this tendency,” said David Brent of the University of Pittsburgh, who participated in the study.
According to fellow researchers, their method is interesting, but not very practical. After all, in order to distinguish a potential suicide from a normal person, doctors will have to put the patient in a scanner, make a tomogram, after which it will be possible to understand whether this person wants to kill himself or not. Yes, and scanners of this kind are far from being in all hospitals, but only in specialized ones.
In order to make their methodology more practical, its authors
plan to use encephalograms to identify potential suicides. Systems for removing EEGs are much more portable, they are in almost every hospital, and their use is less problematic than working with scanners.
Whatever it was, but the work of doctors is very interesting - for the first time someone was able to show exactly how a person who has suicidal tendencies differs from an ordinary person.

